INTRODUCTION
Cell is the structural and functional unit of life. Human body is made of various types of cells. The fundamental characteristics of these cells are essentially same. However, during the process of differentiation, the cell acquires many unique morphological and functional properties.
EUKARYOTIC VERSUS PROKARYOTIC CELLS
The cells are mainly classified as prokaryotic and eukaryotic cell (Table 1.1). The cells of bacterial and other lower organisms are known as prokaryotic cell. The higher animals are made of eukaryotic cells. In prokaryotic cell, deoxyribonucleic acid (DNA) is present within the cytoplasm without any distinct nucleus. The main distinguishing features of eukaryotic cells are:
- Deoxyribonucleic acid is enclosed by membrane.
- The cell contains mitochondria and other membrane bound vesicles (Table 1.1).
- Deoxyribonucleic acid is double stranded compared to circular DNA in prokaryotic cells.
Unlike histology, in cytological examination detailed cell-to-cell relation is often lost.
Cytologist studies cluster of cells or single cell for diagnosis. It is essential to know the detailed morphology and function of the cell to understand the alteration of its constituents in reaction to various external and internal stimuli. The various morphological constituents of a cell are highlighted in Box 1.1 and demonstrated in Figure 1.1. Electron microscopic features of the cell are shown in Figure 1.2. The constituents of the cell can be divided into cytoplasmic organelles and nucleus.
Cell Membrane (Fig. 1.3)
Cell membrane is a selectively permeable biological membrane that separates the cell from its interior and the external environment.
Singer SJ and Nicolson GL1 first proposed the “fluid mosaic model”. They said that the plasma membrane is just like fluid. The membrane proteins are floating on discontinuous fluid like lipid bilayers. The proteins of the membrane are a set of heterogeneous globular molecules. The highly polar groups are protruding out of the membrane and the nonpolar groups are within the interior portion of the phospholipid membrane. The membrane is described as “mosaic” because it is composed of different types of molecules such as phospholipids, glycolipids, cholesterol and proteins.
Composition and Structure (Box 1.2)
The cell membrane is composed predominantly of bilayered phospholipid molecules, proteins, and carbohydrates.
Lipids: There are three classes of lipids: (1) phospholipid, (2) cholesterol, and (3) glycolipid. The phospholipids are the predominant type of lipids noted in the cell membrane. There are four varieties of lipids: (1) phosphatidylcholine, (2) phosphatidylethanolamine, (3) phosphatidylserine, and (4) sphingomyelin. Phospholipids have the hydrophilic or polar ends and hydrophobic or nonpolar ends. In the hydrophilic ends usually the glycerol molecules combine with serine, choline or ethanolamine, whereas, in the hydrophobic ends the glycerol molecule is attached with the long chain fatty acids.
Hydrophobic ends of the molecules are facing each other and they are away from the cytosol or external environment. Whereas the hydrophilic ends are facing toward the cytosol. At low temperature, the bilayered lipid is just like gel. However, in body temperature, the lipid bilayer is fluid and moving and can exchange their places.
Good amount of cholesterol molecules are also present in the plasma membrane and one cholesterol molecule is present for one phospholipid molecule. The cholesterol molecules are embedded within the phospholipid layers. They prevent the mobility of the first few hydrocarbon molecules of the phospholipid and also prevent the crystallization of the hydrocarbon. Thus, cholesterol maintains the fluidity and stability of the membrane.5
Carbohydrate: Carbohydrates are present in the form of glycoprotein and glycolipids. Glycoprotein is the predominant type of carbohydrate and is generally noted on both sides of the membrane. They are involved with cell recognition and protection of the membrane.
Proteins: Proteins are the 50% constituents of the membrane and depending of their positions they may be labeled as:
- Integral protein: Integral proteins are incorporated within the membrane (Fig. 1.3). Transmembrane proteins are the type of integral proteins that traverse through the complete breadth of the membrane.
- Peripheral membrane protein: These proteins remain only on the inner or outer surface of the membrane.
Functions of Plasma Membrane (Box 1.3)
The plasma membrane is biologically active semi-permeable membrane with many important functions.
- Cell identity: Plasma membrane encircles the essential component of the cell and acts as a physical barrier between the cell and its surroundings.
- Transport: Plasma membrane is selectively permeable to various substances. As the membrane is hydrophobic in its interior so it is impermeable to most polar molecules such as Na+, H+ and Cl-. However, the lipid bilayer is permeable to small non polar molecules such as CO2 and O2. There are two types of transport through the membrane:
- Active transport: Energy is needed for this type of transport.
- Passive transport: No additional energy is needed. This can be channel or transporter protein mediated facilitated diffusion or by osmosis. In osmosis passive transport occurs across the concentration gradient. In case of facilitated diffusion, two types of protein take part in action:
- Transporter proteins: This type of protein alters the conformation of the solutes to be transported and sequentially transport the solute through the lipid bilayer.
- Channel protein: This protein forms an aqueous pore across the membrane.
- Signal transduction: Plasma membrane contains many membrane-bound receptors. These receptors bind with signaling molecules and transport the information via the intracellular signaling proteins. The signaling molecules may be soluble, attached with the other cell or may be bound to the extracellular matrix. The three major classes of plasma membrane receptor proteins are involved in signal induction (Fig. 1.4):
- Ion-channel-coupled receptors: These receptor proteins are involved in transient opening and closing the ion channel after binding with the signaling molecules.
- G-protein-coupled receptors (GPCR):2 The GPCRs mediate their actions by transiently binding with a trimeric GTP (Guanosine triphosphate)-binding protein (G protein) (Fig. 1.4).
- Enzyme-coupled receptors: These receptor proteins are predominantly protein kinases. In their activated form they phosphorylate the specific types of proteins.
- Cell polarity (Box 1.4): Most of the cells in human body are polarized. Cell polarization is studied in the epithelial cell. The epithelial cells have distinct polar distribution such as luminal surface and basolateral surface facing towards the basement membrane and side of the cell. Various membrane protein complexes are responsible for the polarity of the epithelial cells. Three types of polarity complex proteins are described in the membrane of the epithelial cells: (1) PAR (CDC42–PAR3–PAR6–aPKC), (2) Crumbs (Crb–PALS–PATJ), and (3) Scribbles (Scrib–Dlg–Lgl). PAR and Crumbs complexes are involved in the apical polarization and Scribbles complexes are responsible for basolateral polarization of the epithelial cells. They are also involved in the asymmetric cell division, cell proliferation and cell migration.3 Asymmetric cell division suppresses cell proliferation. Disruption of the polarity complexes is related with cell proliferation. It is noted that the loss of epithelial cell polarity complexes is related with tumor, progression and invasion.4 So, in fact, the membrane polarity complexes behave as tumor suppressor elements.
Epithelial Mesenchymal Transition and Cell Polarity (Box 1.5)
Epithelial mesenchymal transition (EMT) is an orchestrated series of events in which the epithelial cells changes by shedding out some characteristics properties and acquiring many typical properties of the mesenchymal cells. It is one of the key steps during embryonic development, chronic degenerative fibrosis, and cancer metastasis.5 During the process of EMT in vitro, the following steps happens (Fig. 1.5):
- The epithelial cells lose polarity
- Separation of the individual cells, and
- Gaining of cell motility followed by subsequently dispersion of the cells.6
During intravasation, tumor cells pass through the endothelial membrane of the vascular channels and come into the systemic circulation. In extravasation the tumor cells attach to the blood vessels and leave the bloodstream.7 Finally the tumor cells make new colony in the new host environment. It is very much essential to understand the initial steps of metastasis because this knowledge may help to develop strategies to prevent metastasis of tumor cells. EMT plays significant role in tumor metastasis.
Fig. 1.5: Schematic diagram showing epithelial mesenchymal transition related with metastasis.
(MMPase: Matrix metalloproteinase).
7As mentioned before, the typical epithelium is sheet of cells where the cells are attached with the basal basement membrane and laterally with other by cell junctions. These cell junctions prevent the movement of the individual cells. Mesenchymal cells are usually more elongated and they show front to back polarity rather than apicobasal polarity. In EMT apical/basolateral polarity of the epithelial cells are lost due to decreased expression of cell polarity proteins.7 Intracellular architecture of the epithelial cells is also lost so that the cells gain mobility.
E-cadherin, a transmembrane protein, helps to maintain the adherens junctions. E-cadherin molecules bind cytosolic catenin with the help of its intracellular domain and make a link with actin cytoskeleton. The extracellular domain of this molecule helps in the interaction with other neighboring cells. The epithelial cells become loose in attachment and become movable when they lose E-cadherin from their membrane.
In EMT, the transformed epithelial cells show increased expression of:
- N-cadherin
- Vimentin
- Catenin
- Matrix metalloproteinases.8
N-cadherin is usually present in the mesenchymal and is not normally expressed in the epithelial tissue. The increased expression of N-cadherin in the epithelial cells facilitates their movement within the stromal tissue.8 The transformed cells in EMT also produce increased amount of transcription factors such as Snail 1 and Snail 2, vimentin, catenin and matrix metaloproteinase.8 Overall, due to EMT the transformed cells acquire the properties of increased migration, invasion and scattering.
- Cell-to-cell recognition: Glycolipids and glycoproteins are responsible for mutual cell-to-cell recognition.
- Intercellular joining: One of the important functions of the plasma membrane is connection between two cells. This is discussed below.
- Attachment to the cytoskeleton and extracellular matrix.
Cilia and Flagella
Cilia and flagella are the mobile extension from the surface of the cytoplasm (Box 1.6). Cilia are the small, regular and multiple in number, whereas, the flagellum is single slender structure. Cilia are seen on the lining epithelium of the upper respiratory tract and fallopian tube. Each cilium is attached with the thick terminal plate near the apical surface of the cell. The ciliated cells are usually polar and are attached with the basement membrane. The central elongated portion of the cilium is known as axoneme. At the base of the cilium or flagellum is a basal body. The basal body is composed of microtubules. There are total 11 microtubules. In the center there are two singlet microtubules surrounded by nine triplet microtubules (Fig. 1.6). The outer peripheral doublet microtubules in the axoneme have a pair of dynein arms that are extended to the adjacent microtubules. These dynein arms help in the movement of the cilium and flagellum. Cilia are usually lost in the cancer cell originated from the bronchial epithelium. Therefore, the presence of cilia on the cell almost safely excludes the possibility of malignancy.
Function: Cilia help in the movement of the particles or organism in one direction.
Fig. 1.6: The structure of cilia is shown. The long shaft of axoneme is originated from the basal body. The cross-section of axoneme shows 9 doublet microtubules and two central singlet tubules.
Brush Border
The surface of the certain specialized epithelial cells covered with multiple microvilli is known as brush border (Box 1.6). They are regular finger-like projections on the cell surface about 1 micron in length. Microvilli are commonly seen on the luminal surface of the intestinal epithelium (Fig. 1.7) and also on the proximal tubular epithelial cells of kidney. On light microscopy the microvilli are seen as fuzzy appearance.
Function: The brush border increases the surface area of the cell and helps in better absorption of the substances from the large surface area.
Cell Junction
Cell junction can be classified by depending on localization of the junction (Fig. 1.8):
- Cell to cell:
- Tight junction
- Adherens junction
- Desmosomes
- Gap junction
- Cell to matrix: Hemidesmosomes
- Tight junction: This is located in the apical region of the epithelial cells and almost completely seals the gap between the two epithelial cells toward the luminal site. The sealing strands of transmembrane adhesion proteins encircle the apical part of the cell membrane of the two cells and hold them tightly. Claudins and occludins are two major transmembrane adhesion proteins.Fig. 1.7: Electron microscopic picture of brush border of intestinal cell.Courtesy: Dr Charan Singh Rayat, Department of Histopathology, PGIMER, Chandigarh, India.
- Function: There are two “major functions” of the tight junctions: (1) Tight junction closes the gap between luminal side and the intercellular space. This helps in effective transport of substances from luminal side of the cell to extracellular fluid compartment. (2) It prevents the drift of the apical membrane proteins to the basal region and vice versa.
- Adherens junction: The adherens junction holds the two cells together and confers mechanical strength. Adherens junction is made of cadherin, catenin and intracytoplasmic actin filaments. Altogether they form adhesion belt like structure.
- Function: To provide cell-to-cell adhesion and mechanical strength.
- Desmosomes: These are button like spots which connect the plasma membrane of two cells together. Desmosomes are linked to the intermediate filaments. The type of intermediate filament depends on type of the cell.
- Function: The desmosomal junction provides tensile strength and rigidity of the tissue.
- Gap junctions: These are intercellular channels that connect two adjacent cells. In gap junctions, the two plasma membranes are connected by the transmembrane proteins known as connexins.
- Function: There are continuous channels between the two adjacent cells and therefore the cells can rapidly share small molecules and ions. With the help of the gap junctions the action potential can rapidly travel among a group of cells without any neurotransmitter.
- Hemidesmosomes: Hemidesmosomes connect the cell with the basal lamina. Hemidesmosomes are composed of keratin filaments, dystonin, plectin, integrin, collagen XVII and laminin. In hemidesmosome, integrin binds with keratin in cytoplasmic side by dystonin and plectin. It also binds with collagen XVII and laminin toward basal lamina side.
- Function: It attaches cell with the basal lamina.
Cytoplasmic Organelles
These are tubular and cistern like spaces and vesicular structures folded within the cytoplasm. Endoplasmic reticulum (ER) is generally connected with cell membrane to nuclear membrane (Box 1.7). The cisterns are membrane like long flat spaces which are straight, whereas tubules are irregularly branched structures. The ER contains fluid with many enzymes and proteins. Two types of ER are present:
- Rough endoplasmic reticulum (RER), and
- Smooth endoplasmic reticulum (SER).
Rough endoplasmic reticulum: They are tightly packed parallel bundles of cistern like spaces which are beaded in appearance due to ribosome particles attached to the surface of RER. The ribosomes are bound with RER by a receptor known as “Rhiboporin”.
Fig. 1.10: Electron microscopic picture of rough endoplasmic reticulum.Courtesy: Dr Uma Nahar Saikia, Professor, Department of Histopathology, PGIMER, Chandigarh, India.
Function: It is the site of synthesis of secretory protein and lysosomal enzymes.
Smooth endoplasmic reticulum: The SER predominantly contains tubules and vesicles. SER is connected with Golgi apparatus and plasma membrane.
Golgi Complex (Box 1.8)
Golgi apparatus are stacks of membrane bound cistern like spaces within the cytoplasm arranged in polarized fashion. Each stack of Golgi complex (GC) has four parts (Fig. 1.9):
- 1. Cis-Golgi network: Cis-Golgi network is the concave surface of the stack of GC that faces toward the RER and small transfer vesicles. Cis-Golgi network receives the initial protein from ER.
- 2. and 3. Endo-Golgi and Medial-Golgi network: These are the middle parts of GC and most of the proteins are modified here.
- 4. Trans-Golgi network: This is the convex surface of the stack of the GC. Trans-Golgi network is associated with large secretory vesicles and final transport of the protein.
Function: The main function of the GC is chemical processing of the protein received from RER followed by packaging and transfer. Along with classical “protein trafficking”, there are many other novel functions of GC such as entry of the cell to mitotic check point, calcium homeostasis and cytoskeletal organization.10
Protein modification: N-linked and O-linked glycosylation of protein and lipids occur in the GC.
Protein transport: Golgi complex receives the neosynthesized protein from the ER and transport to their respective destination. The cargo proteins are first modified, and then they are transported by GC. The mechanism of the transport of cargo proteins is not exactly known. However, there are two theories:
- Vesicular transport model theory: The cargo protein is transported by an anterograde way with the help of vesicles that bud from one cisterna and then fuse to the next one.
- Cisterna maturation model: In this model, it is assumed that the Golgi cisterns are formed de novo, progressively mature and finally dissipate.11
Calcium storage: Golgi complex is the most important site of intracellular calcium storage and can also release Ca2+ in case of agonistic stimulation.
Platform of different cells signaling: Golgi complex acts as the platform of different signaling events within the cell. In addition to the receiving initial signal, GC can also induce a cascade of signal transduction.
Mitochondria
These are 0.5–1 µ diameter organelle considered as the power house of the cell as they are the major source of energy (Box 1.9). Mitochondria (MT) are richly localized in that part of the cells that requires energy such as interfibrillar space in the striated muscle and in the middle part of the sperm. Other than nuclear DNA, mitochondria have its own independent DNA and this is the unique feature of MT.
Structure
Mitochondria are double membrane-bound structures, which consist of following parts (Figs. 1.11 and 1.12).
- Outer membrane
- Inner membrane
- Intermembranous space
- Cristae
- Mitochondrial matrix
Outer membrane: This is composed of phospholipid bilayers. The outer membrane of MT is rich in porin, a variety of integral proteins. This forms a through and through aqueous channel in the outer membrane. Therefore, the small ions and proteins can easily cross the outer membrane. MT associated endoplasmic reticulum connects the ER and outer membrane of MT.
Inner membrane: This is the inner phospholipid bilayer of the MT. The inner membrane is rich in double phospholipid known as cardiolipin. The cardiolipin possesses four fatty acids rather than two, and the presence of the cardiolipin makes the inner membrane impermeable to proton, ion and electrons. In certain regions the outer and inner membrane joins together known as contact sites and makes a passage of the proteins and small molecules from the cytoplasm to the matrix space. The inner membrane contains the large number of lollypop like structure with small stalk attached to the inner membrane and globular region in the matrix. These globular regions contain protein complex of ATP (adenosine triphosphate) synthase. The inner membrane contains three types of enzyme: (1) ATP synthase, (2) the respiratory chain protein complexes such as NADH dehydrogenase complexes, cytochrome b-c1 and cytochrome oxidase, and (3) transport protein complexes.
Intermembranous space: This is the tiny space bounded by two membranes of the MT. The concentration of small molecules is same in both cytoplasm and intermembranous space.
Cristae: this is the small shelf like folds of the inner membrane. This makes the larger space in the inner membrane to retain more enzymes.
Matrix: It is the innermost space of MT encircled by the inner membrane. The matrix consists of dense fluid that is rich in viscosity. Matrix is rich in enzymes of citric acid cycle and also contains mitochondrial DNA.
Fig. 1.12: Electron microscopic picture of mitochondria.Courtesy: Dr Uma Nahar, Saikia, Professor, Department of Histopathology, PGIMER, Chandigarh, India.
Functions
Main function of MT is energy production in the form of ATP synthesis. However, it is also involved in other important functions such as calcium storage, cell death, etc.12
- Citric acid cycle: The essential enzymes in citric acid cycles are located in the mitochondrial matrix, and the main reactions of citric acid cycle happen in the mitochondrial matrix. Initial oxidative breakdown of glucose occurs in the cytoplasm by the process known as glycolysis.
- Electron transport: During the oxidative phosphorylation, a series of electron transport reaction occurs in the inner mitochondrial membrane. The energy released in these reactions is used to generate ATP from ADP.
- Calcium storage: Mitochondria can store calcium and play an important role in calcium homeostasis.
- Cell cycle: Various signaling reactions of cell cycle occur in the MT.13
Mitochondrial Deoxyribonucleic Acid
Mitochondrial DNA is unique to MT and it is considered as separately developed during evolution (Box 1.10). In sexual reproduction, MT is exclusively inherited from mother and so MT DNA is of maternal origin. MT DNA is organized in nucleoid. The nucleoid is the combination of MT DNA and proteins. The nucleoid is also known as DNA-protein complex (DNP). MT DNA is 16.6 KB circular DNA molecule that encodes only 37 genes. It is responsible for the production of selected MT protein, ribosomal RNA (rRNA) and transfer RNA (tRNA). The remaining 99% proteins in the MT are produced by nuclear MT gene. Therefore two parallel encoding of proteins in MT are going on (1) by MT DNA and (2) Nuclear mitochondrial DNA. Replication of mitochondrial DNA is dependent with the cell cycle.
Mitochondrial dysfunction may occur due to defects in MT DNA or nuclear mitochondrial DNA (nDNA). Interestingly, mutations in the MT DNA are heteroplasmic that means both mutant and wild type molecules remain in the same cell in varying proportion. The cell can tolerate a threshold of defective proteins and after that behaves abnormally. In course of time all MT DNA may be of same genomic type. This is known as homoplasmy. Two types of mutation may occur in MT DNA: (1) point mutation, (2) deletion.15
Point mutation: Almost all the genes of MT DNA are affected by point mutation. This is maternally transmitted. The mother may be unaffected due to less than critical threshold to produce the disease. However, the child may be affected by the disease. The patient may have Mitochondrial Encephalopathy, Lactic Acidosis and Stroke (MELAS).
Single large scale deletion of MT DNA: The deletion of MT DNA is usually sporadic and occurs de novo. It involves several genes. The diseases produced by deletion of MT DNA include progressive external ophthalmoplegia, Pearson syndrome and Kearns Sayre syndrome.16
Nuclear DNA mutation causing mitochondrial disease: These diseases follow Mendelian inheritance and they are X-linked (both dominant and recessive). De novo mutation may also occur. The mutation of nDNA genes may cause:
- Defects in structural subunit of oxidative phosphorylation complexes
- Defects in mitochondrial import (dilated cardiomyopathy, spastic paraplegia, etc.)
- Defects in mitochondrial fusion (Charcot-Marie-Tooth disease)
- Defects in mitochondrial translation
- Defects in mitochondrial maintenance (myopathy, encephalomyopathy, etc.)
- Coenzyme Q10 deficiency
- Defects in F-S clusters (Friedreich-ataxia syndrome, multiple mitochondrial dysfunction syndrome).
Ribosomes
Ribosome is the sites of protein synthesis (Box 1.11).
Structure (Fig. 1.13)
Ribosomes are small 25–30 nanometer particles present in the cytoplasm. They are present both as free ribosomes in cytosol and also in the membrane-bound form attached with ER and thus forming rough endoplasmic reticulum. The ribosome is made up of r-RNA and proteins. The ribosome is classified according to the sedimentation coefficient in ultracentrifugation. The eukaryotic ribosome has two units a smaller 40s and a larger 60s subunit. In its inactivated form the two subunits are detached however when the ribosome is engaged in protein synthesis both the units are attached together. Small subunit has the binding sites for mRNA and t-RNA. Some rRNA of the larger subunit has enzymatic activity to catalyze the peptide bond. These rRNA are known as the ribozyme.
Function
Ribosomes take part vital role in protein synthesis by decoding information from mRNA and then help molecules of tRNA to assemble particular amino acids to make a protein.
Lysosomes
These are 0.2–0.4 µ small membrane-bound vesicles present in the cytoplasm (Box 1.12). They contain near about 40 acid hydrolytic enzymes. Lysosomal enzymes are produced in the endoplasmic reticulum and carried over in Golgi complex. The lysosomal enzymes and membrane of the lysosome are finally synthesized in the trans-Golgi network and are carried to the endosome by clathrin coated transport vesicles. The final lysosomal vesicles are synthesized in the late endosomal intermediate (also known as “endolysosome”).
There are two types of lysosome: (1) Primary lysosome: No morphological sign of hydrolytic enzymes, and (2) Secondary lysosomes: When lysosome fuses with other phagocytic vesicles of an organism it shows enzymatic activities and this lysosome is known as “secondary lysosomes”.
Functions
Lysosome contains acid hydrolytic enzymes such as (1) lipase, (2) amylase, (3) protease, and (4) nuclease. These enzymes are activated in the acid environment. The lysosomal enzymes digest the macromolecules, destroy the microbes and remove the other cytoplasmic organelle such as mitochondria. The foreign organisms enter the cytoplasm as phagocytic vesicles. Lysosome fuses with the phagosome and release acid hydrolytic enzymes, which degrade the protein and carbohydrate components of the organism. The lipid component is more resistant to digestion and may remain as the residual body. At times, lysosome fuses with the nonfunctioning MT or fragments of RER to clear these substances from the “autophagic” vacuoles in the cytoplasm. When these autophagic vacuoles remain persistently in the cytoplasm, they accumulate pigment known as “Lipofuscin”.
Peroxisome
These are tiny vesicles of 0.2–1 µ in size. They are synthesized from RER. Peroxisomes contain many oxidative enzymes. The enzymes in the peroxisome break down fatty acid by beta oxidation and generate acetyl coenzyme A and H2O2. Acetyl coenzyme A is involved in various energy producing metabolic processes. The hydrogen peroxide helps to kill the various organisms. Excess hydrogen peroxide is further degraded by catalase enzyme of peroxisome into water and oxygen.14
Cytoskeleton (Box 1.13)
Cytoskeleton is the meshwork of protein filaments within the cytoplasm that maintains the shape of the cell along with other important functions such as cell movement, cell contraction and maintaining cell polarity. There are three components of the cytoskeleton:
- Microfilament
- Microtubules
- Intermediate filaments
Microfilament
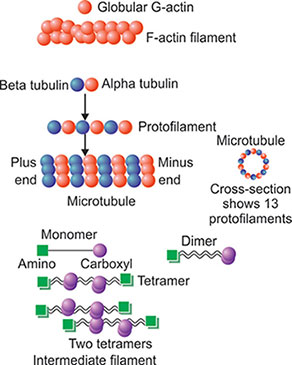
It is also known as “actin filament”. These are the thinnest filaments and present either as a bundle form or as fine network within the cytoplasm. The actin filament is composed of globular G-actin which is polymerized and forms a long chain of F-actin (Fig. 1.14). Most of the G-actin is bound with small proteins such as profilin and thymosin. Binding of these small proteins prevents the polymerization of G-actin. Actin filament binds with filamin and makes a robust supporting meshwork just underneath the plasma membrane known as “cell cortex”. There are three types of actin: (1) Alpha actin, (2) Beta actin, and (3) Gamma actin. Alpha actin is present in muscle and other two forms of actin are present in nonmuscular cells.
Fig. 1.14: Three types of cytoskeletal structures: Actin filament, microtubules and intermediate filament.
Functions
- Actin maintains shape of the cell.
- Actin binds to the myosin and helps in contraction of muscle fibers.
- Actin can shorten its length and helps in the movement of the cell.
- Phagocytosis or pinocytosis is helped by actin.
- Actin helps in the transport of various vesicles within the cytoplasm.
Microtubules
Microtubule is long, straight, hollow rigid tubules of 25 nanometers in diameter. These are dynamic fibers that mean they are always in the process of assembling and disassembling. The microtubules constitute mitotic spindles, centrioles, cilia and flagella. The basic constituents of the microtubules are α- and β-tubulin.
These tubulins are arranged alternatively to form a protofilament (Fig. 1.14). GTP is tightly bound with α tubulin and resistant to hydrolysis whereas it is loosely bound with β tubulin and can be separable by hydrolysis. The protofilament of tubulin is polar as one end, is formed by β tubulin and other end is formed by α tubulin. The β 15end of the tubulin protofilament is plus end as the growth and shrinkage of this end is rapid. The opposite α tubulin end is known as the minus end. In each microfilament, there are 13 total protofilament attached parallel with each other with a central hollow structure. All the plus ends or growing ends of the protofilaments are in one direction.
Functions: The main functions of the microtubule are:
- Intracellular transport: Microtubules help in the transport of the vesicles containing proteins from the Golgi complex to plasma membrane.
- Mitotic spindle movement: The mitotic spindles are formed by microtubules. The chromatids are separated and pulled to each daughter cell nucleus by the mitotic spindles formed by microtubules.
- Movements: Movements of cilia and flagella are done by the microtubules.
Centrosome
It is small round body located near the nucleus in the interphase cell. This is also known as microtubule organizing center. The microtubules are attached with the centrosome by their minus ends, and they radiate from the centrosome like a star-shaped manner. Centrosome consists of a pair of centrioles arranged in L-shaped manner surrounded by the amorphous matrix material known as centrosome matrix or pericentriolar material. Centrosome matrix material takes the main role in the development of the microtubule. Centrioles are the basal bodies of cilia or flagella. During mitosis, the centrosome duplicates and each one contains one pair of centrioles. From each of the centrosomes, microtubules radiate and form a complete mitotic spindle.
Intermediate Filament
Intermediate filaments have average diameter 10 nanometers. The name of the intermediate filaments is such because the diameter of intermediate filaments is in between the microfilament (7 nanometers) and microtubules (25 nanometers). The intermediate filaments are made of individual polypeptides. These are alpha helix structures that contain 310–350 amino acids. It has N and C terminals. Two alpha helix monomer coils with each other to form a dimer. Both the N and C terminals are in same direction in this monomer. Two dimmers then coil in a staggered anti-parallel fashion to form a tetramer. Eight such tetramers twist in a rope like manner to form an intermediate filament. Therefore, in a cross-section of intermediate filament there are 32 alpha helix coils.
- Type I (acidic) and type II (basic): Present in epithelial and hair keratins.
- Type III:
- Vimentin: Vimentin is expressed predominantly in mesenchymal cells and also in leukocytes, vascular endothelial cells and some epithelial cells.
- Desmin: Present in the skeletal and cardiac muscles.
Table 1.2 Intermediate filaments. TypeVarietiesLocationMolecular weight (Da)FunctionIAcidic keratin (11 epithelial keratin, four hair keratin)Epithelial cells40,000–70,000Tensile strengthIIBasic keratin (8 epithelial keratin and 4 hair keratin)Cells of hair and nail40,000–70,000Tensile strengthIII- Vimentin
- Desmin
- Glial fibrillary acidic protein
- Peripherin
- Mesenchymal cells, leukocytes, vascular endothelial cells and some epithelial cells
- Skeletal and cardiac muscle fibers
- Astrocytes and other glial cells.
- Neurons of the dorsal root ganglia, sympathetic ganglia and cranial nerves
- 54,000
- 53,000
- 50,000
- 56,000
- Support the cytoplasmic membrane and helps in holding the various organelles in proper position.
- Helps in stabilizing sarcomeres of the contracting muscle cells
- Supports the glial cells
- Supports the neurons
IVNeurofilaments (NF)- NF-Light
- NF-Medium
- NF-High
Mature neurons- 62,000
- 102,000
- 110,000
They form the cytoskeleton of dendrites and axons.VLamins A, B and CNuclear envelope65,000–75,000Control of assembly of the nuclear envelope during mitotic event and chromatin organizationVINestinStem cells of the central nervous system200,000 - Peripherin: It is present in the peripheral neurons and cranial nerves.
- Type IV: Neurofilaments: Neurofilaments are expressed in the mature neurons.
- Alpha internexin: They are present in developing central nervous system.
- Type V: Lamin: They are present underneath the nuclear membrane (Lamin A, Lamin B and Lamin C). Lamin helps in chromatin organization and gene expression.
- Type VI: Nestin: Nestin is present in stem cells of the central nervous system and in developing striated muscle.
Unclassified
Filensin: It is present in the epithelial cells of the lens at the time of differentiation.
Functions of the Intermediate Filaments
- Supporting the cytoskeleton structure: Intermediate filaments are more stable than microtubules and microfilaments and provide good support and tensile strength of the cytoskeleton of the cell. Desmin links myofibrils of the striated muscles. GFAP supports the glial structure, and neurofilaments support the cytoskeleton of the axons and dendrites.
- Chromatin organization: Nuclear lamin takes part and plays an important role in chromatin organization of the nucleus. Lamin also controls the assembly of the nuclear envelope at the time of mitosis.
Nucleus
Nucleus is the central processing unit of the cell and act as the controlling center of the cell (Box 1.14). The important components of the nucleus are:
- Nuclear envelope and pore
- Nuclear matrix
- Nuclear chromatin
- Nucleoli.
Nuclear Envelope
Nuclear envelope is the barrier between the nucleus and cytoplasm. It has three parts:
- Nuclear membrane
- Nuclear pore
- Nuclear lamina.
Nuclear Membrane (Figs. 1.15 and 1.16)
The nuclear membrane is further divided into (a) outer nuclear membrane (ONM), (b) inner nuclear membrane (INM) and (c) perinuclear space.
The ONM is the most outer part of the nuclear membrane, and it is 6 nm thick. ONM is continuous with the ER. The ONM is usually studded with multiple ribosomes on its cytoplasmic side that are involved in protein synthesis. The INM is parallel to ONM and is directly attached to the nuclear lamina. The space in between the ONM and INM is known as perinuclear space. The width of this space is 50 nm. Both the ONM and INM are perforated by multiple nuclear pores.
The nuclear lamina is situated in the inner side of the INM. It is made up of nuclear lamin that is intimately related to the cytoplasmic intermediate filaments. The INM contains lamin B receptor (LBR), lamina associated polypeptide (LAP), MAN 1, emerin, nurim and ring finger binding protein (RFBP).18
Functions
- Acts as a physical barrier between the cytoplasm and nucleus
Fig. 1.16: Electron microscopic picture of nucleus and its membrane.Courtesy: Dr Uma Nahar Saikia, Professor, Department of Histopathology, PGIMER, Chandigarh, India.
- Various integral proteins such as LBR, LAP, RFBP help in chromatin remodeling and gene expression. LAP and lamin A/C bind with Rb-protein that further recruits histone deacetylases (HDAC), DNA methyl transferases (Dnmt 1), histone methyltransferases (HMTase) and heterochromatin protein 1 (HP1). The action of these enzymes changes the higher-order conformation of chromatin, and ultimately causes gene silencing by inhibiting transcriptional activation of E2F.19
Nuclear Pore
At certain positions: the ONM and INM fuse with each other and, therefore, make the pore on the nuclear membrane that makes the direct connection site between the nucleus and cytoplasm. The diameter of each nuclear pore is about 100 nanometer. The number of nuclear pore varies from few 100s to 1,000s depending on the metabolic activity of the cell. The nuclear pore complex (NPC) is the gateway of the nucleus across the double membrane nuclear envelope (Fig. 1.16) that selectively allows the passage of various macromolecules across the nucleus.19
Nuclear pore complex consists of a cytoplasmic ring, a nuclear ring and a distal ring connected by nuclear basket (Fig. 1.15).20
Function: The main function of the nuclear pore is the facilitation of the cytoplasmic to nuclear traffic and vice versa.
Nuclear Matrix
It is the internal skeleton of the nucleus and consists of an RNA network, protein complexes, peripheral nuclear lamin and residual nucleoli. Nuclear matrix is the remaining part of the nucleus after removal of chromatin, nuclear membrane and the other soluble components of the nucleus. Its composition is dynamic and varies with nuclear activities. In fact, nuclear matrix protein (NMP) may be just a processional artifact only as its demonstration depends on the selected conditions on removing the chromatin and lamin. The NMP is tissue specific and it differs from normal and neoplastic tissue of the same type. NMP is interlinked with the intermediate filaments of the cell cytoplasm and provides the overall support of the nucleus. Nuclear matrix contains both proteinaceous substances, nuclear mitotic apparatus protein, actin and RNA.21 The chromatin forms a loop and the base of the loop is attached with nuclear matrix. The strings of DNA attached with the nuclear matrix is labeled as matrix attachment region (MAR) and the corresponding nuclear matrix is called MAR-binding proteins. Active sites of genes are usually located in MAR sites than the loop areas. Other than gene transcription, NMP also participates in gene translation.
Nuclear Chromatin
Chromatin Structure
Chromatin represents the uncoiled chromosome of the interphase nucleus (Box 1.15). It is composed of DNA, histone and nonhistone proteins.22 In the interphase, the individual chromosomes are present in the specific region of the nucleus which is known as chromosomal territories. The channels that separate the chromosomes are known as interchromosomal domains.
Chromatin can be classified as heterochromatin and euchromatin (Table 1.3).
Heterochromatin
Heterochromatin is located on the nuclear membrane (Fig. 1.2). It is the condensed part of chromatin and contains inactive genes.18
|
Heterochromatin is further divided into facultative heterochromatin and constitutive heterochromatin. The facultative heterochromatin contains inactive genes in certain stages of development of the cells. The constitutive heterochromatin is made of inactive genes that are the structural components of the chromosomes, e.g. telomeres and centromeres.
Euchromatin
It is located in the interior of the nucleus. In comparison to heterochromatin, euchromatin is less condensed and consists of the actively transcribed genes.
Nucleosome—the Basic Unit of Chromatin
Nucleosome is the basic unit of chromatin. It is made of approximately 146 bp of DNA. This DNA encircles in two turns around a central octameric protein core. This core histone complex is composed of two copies each of histone H2A, H2B, H3, and H4 (Fig. 1.17). Two copies of H3 and H4 remains in the center as tetramer and two H2A-H2B dimmers located on either side of the tetramer. Therefore, the linear arrangement of these four histones is: (H2A/H2B)-(H4/H3)-(H3/H4)-(H2B/H2A). The DNA chain in between the two nucleosomes is the linker DNA. The linker histone (Histone H1) binds the linker DNA chain and the core histone. The strings of linked nucleosomes are repeatedly twisted to form the more complex threedimensional organization of chromatin.
Histone Modification
The core histones are the dynamic structures and are continuously modified (Fig. 1.18). The tail parts of histone are projected out of the nucleosomes. The various histone modifiers act on the amino terminal ends of these histone tails and control the remodeling of the chromatin structure. The unwound remodeled chromatin facilitates the transcriptional proteins to interact with promoter sequence of DNA and thereby helps in DNA transcription.
Histone Tails Modification22
Acetylation: Lysine side chain of H3 and H4 are acetylated with the help of histone acetyltransferases (HATs) enzyme. This acetylation neutralizes the charge between lysine (positively charged) and DNA (negatively charged). The chromatin becomes opened up to recruit various transcription factors for transcriptional activation.
Histone deacetylase (HDAC): HDAC enzyme helps in deacetylation of histone tails that causes chromosomal recondensation and subsequently repression of DNA transcription.
Histone methylation: Histone methyltransferases (HMT) enzymes help in the methylation of lysine or arginine residues of histone H3 and H4.19
Phosphorylation: Aurora A, B and C phosphokinase help in mitotic phosphorylation of histone H3 and take important role in transcription, DNA repair and apoptosis.
In addition, histones are also modified by ubiquitylation and sumoylation.
Histone code: Histone code is a hypothesis that says that DNA transcription is regulated by the modification of histone proteins. It hypothesizes that there may be a particular set of combination of modification of histone tail. This pattern may produce the code which is read by the particular histone modifiers. There may be bromodomain that binds to acetylated lysine causing gene transcription. Whereas, chromodomain specifically binds with methylated lysine causing transcriptional repression.23
Nucleoli
Nucleolus is the subnuclear round to oval small structure within the nucleus and about 1 µ in diameter (Box 1.16). It is not a membrane bound structure. Nucleolus is usually situated in the center of the nucleus; however, the position of the nucleolus may vary. The number of nucleolus may vary from 1 to 3. The size of the nucleolus depends upon the requirement of ribosome and protein synthesis. So it is expected that metabolically active cell with higher amount of protein synthesis will have larger nucleoli. Nucleolus is easily recognizable by light microscopic examination. It looks like eosinophilic round body in H & E smear and light blue colored structure in MGG stain.
The nucleoli are formed at the end of the mitosis (Fig. 1.19A). The specific genetic loci of the origin of nucleoli are known as “nucleolar organizing regions (NORs)”. The nucleolar organizer loci are seen in homologous chromosomes of 13, 14, 15, 21, and 22. Therefore at the end of mitosis, tiny 10 nucleoli appear to from the NOR of the five pair of chromosomes (total 10 chromosomes). These small nucleoli conglomerate to form a single larger nucleolus. The nucleolus contains protein and ribosomal RNA. The protein and r-RNA are surrounded by chromosomal DNA of the nucleolus.
Structure
On electron microscopy, the nucleoli show three major subregions24 (Fig. 1.19B).
- Fibrillar center (FC): Round structures with different size having very low electron opacity.
- Dense fibrillar component (DFC): It is made of densely aggregated fibrils and is located in the peripheral rim of FC.
- Granular components (GC): This is the outermost region and is composed of granules.
These different regions of nucleoli probably indicate the stages of RNA transcription and ribosomal assembly.
Function
The nucleolus is the site of rRNA transcription, processing and ribosomal assembly.
- rRNA transcription: Active rRNA genes are situated in the FCs and DFCs. RNA polymerase I enzyme helps in the transcription of rRNA genes. The primary transcript of the ribosomal DNA is pre-rRNA.20Subsequently, the preribosomal RNA transcripts are handled in the DFC with the help of small nucleolar RNA and other protein processing factors. A series of cleavages occur during the processing of preribosomal RNA to mature rRNA. In addition, considerable amount of methylation of the bases and ribose residues also happens. The various proteins in the nucleolar organizing regions (AgNOR) are demonstrated by the silver precipitation technique.25
- Ribosomal synthesis (Fig. 1.19B): Outside the nucleus, the genes of the ribosomal proteins are transcribed and from the cytoplasm these proteins are transported to the nucleolus. With the help of rRNA these ribosomal proteins are assembled within the nucleolus to form preribosome. The preribosome is transported back to the cytoplasm for final maturation.
Deoxyribonucleic Acid
Deoxyribonucleic acid, the nucleic acid, carries the vital genetic information of the cell (Box 1.17).
The portion of DNA that carries the genetic information is known as “gene”. Within the nucleus DNA is coiled and supercoiled to make a thread like structure known as chromosome. During cell division the chromosomes are visible by light microscopy as a distinct entity and in interphase nucleus the chromosome remain as partly condensed and partly extended form and not possible to locate as a separate entity. There are 23 pairs of chromosomes and out of which 22 pairs are autosomes and one pair is sex chromosome. Sex chromosomes in the male are X and Y chromosomes and in the female are X and X chromosomes. The chromosomes are different from chromatin and the differences are highlighted in Table 1.4.
|
Structure (Fig. 1.20)
Deoxyribonucleic acid is made of double helical strands containing a sugar phosphate backbone and bases attached with the sugar molecule.26 Each strand of DNA is made up of alternate sugar and phosphate molecules. The sugar molecule is a pentose sugar and it is attached with the phosphate by third and fifth carbon atom alternatively. The nucleobase is attached with each sugar molecule and then links with the other base of the opposite strand by a weak hydrogen bond. There are four base pairs: (1) adenine (A), (2) cytosine (C), (3) guanine (G), and (4) thymine (T). Adenine and guanine are purine bases. Cytosine and thymine are pyrimidine bases. Adenine joins only with thymine, and cytosine joins only with guanine. This is known as complementary base pairing. There is another pyrimidine base known as “uracil” that is found in RNA.21
Fig. 1.20: Nuclear chromatin and DNA structure. Double helix DNA structure is made of sugar, phosphate back bone and four bases Adenine (A), Guanine (G), Cytosine (C), and Thymine (T).
Nucleotide
Nucleotide is the basic structural unit of DNA. It consists of a pentose sugar, phosphate and nucleobase (adenine, cytosine, guanine or thymine). Nucleobase and sugar molecule form a nucleoside.
Gene
Gene is the specific portion of DNA with particular arrangement of nucleobases that carries the genetic information for making a particular protein. It is the sequence of A, T, C and G that determine the genetic information. Triplet of three consecutive bases of DNA is known as “codon” and each codon is specific for a particular amino acid. This specifies the sequence of amino acids and subsequently the protein formation. Only the certain parts of the DNA are involved in genetic information to carry and the “in-between part” of the DNA, is commonly known as “noncoding DNA” or “junk DNA”. Near about 98% of human DNA is noncoding and is not involved in protein synthesis. However, it has been noted that the major parts (80%) of DNA are involved in different biochemical activities and therefore are active and are not really junk.27 The exact biological function of the noncoding DNA is not known. Evidences suggest that noncoding DNA interacts with microRNA and thereby controls transcriptional and translation of protein coding sequences.28
Fig. 1.21: DNA replication steps are highlighted in this schematic diagram. Helicase enzyme breaks the double-stranded DNA and a replication fork is formed. With the help of DNA polymerase enzyme DNA strand is made. In the leading strand DNA is synthesized in continuous manner at the direction of replication fork whereas in lagging strand DNA is synthesized in opposite direction as small segments. These small segments are known as Osazaki segments.
Deoxyribonucleic Acid Replication (Fig. 1.21)
Deoxyribonucleic acid replication is a semi-conservative process by which genetic inheritance is maintained. When a cell divides, DNA replication process happens. Here each strand of DNA serves as a template and an identical complementary daughter DNA strand is formed. Each of the newly formed daughter cell contains DNA made up of one original strand and one freshly made strand. Therefore, the DNA double helix replication is a semi-conservative process. This replication process is well controlled and free of mistakes because of stringent proofreading and error checking mechanisms.
Deoxyribonucleic acid replication process needs following enzymes:
Deoxyribonucleic Acid Polymerases
These are the major enzymes in DNA replication process. These enzymes help in the polymerization of 22deoxyribonucleotides into the DNA strand. The DNA polymerase enzyme reads the intact template DNA strand to make the complementary DNA strand. On the basis of sequence homology and structural similarities DNA polymerases are classified into five major families:29 A, B, C, X and Y. The three major varieties of eukaryotic DNA polymerases α, δ and ε belong to family B. The mitochondrial DNA polymerase γ belongs to family A. There are two important and fundamental properties of DNA polymerases:
- They only can add free nucleotides in the 5’ to 3’ direction.
- They need a preformed primer strand which is attached to the template DNA by hydrogen bonding. DNA polymerase adds a deoxyribonucleoside 5’ triphosphate to the 3’ OH group of the primers strand.
DNA replication is the coordinated activities of various enzymatic processes. The basic mechanisms of DNA replication process is evolutionary preserved. DNA replication events are initiated in many hundreds of points in chromosomes. This initiation point of the segment of DNA is known as “origin”. The protein complex that acts on the origin as initiator of the DNA replication is known as “origin recognition complex (ORC)”. The human ORC is site nonspecific. However, it is suggested that in somatic differentiated cells human ORC binds to genomic DNA with certain specificity.30 DNA is replicated in S phase of the cell cycle. Total time of replication of DNA is usually fixed.
For the purpose of description the replication process can be divided by series of steps:
- At first two strands of DNA is separated in particular point known as origin: Here the initiator protein along with other associated protein forms a prereplication complex that separates the two strands of DNA. Therefore, a fork like structure is formed known as “replication fork”. Helicases enzymes break the hydrogen bonds in between the bases and the unwound DNA strands are stabilized by single-stranded DNA binding proteins.
- The binding of RNA primase in the initiation point of the 3’–5’ parent chain: There is extension of the RNA primers by DNA polymerase that binds to the DNA nucleotides of the 3’–5’ strand due to the hydrogen bonds between the bases.
- DNA polymerase adds the matching loose nucleotide: DNA polymerase can act only from 5’ to 3’ direction. Therefore, DNA replication is different in two strands of DNA. Original 5’ to 3’ strand of DNA replication starts from 3’ end and proceeds to the direction of the breakage of the replication fork. The strand of DNA here is known as “leading strand”. In leading strand DNA is synthesized in continuous manner. In other strand of DNA, known as “lagging strand”, the process of DNA synthesis is in discontinuous manner opposite the direction of the replication fork. This occurs in the multiple areas of the DNA strand. Therefore the multiple small pieces of DNA are synthesized known as “Okazaki fragments”.
- Joining of intact lagging strand: The RNA strands are removed by the action of “RNase enzyme” and “DNA Pol I-exonuclease”. The lagging strands are joined by DNA ligase.
Ultimately the DNA replication is terminated. Each double helix DNA contains one old template strand and one newly synthesized fresh strand.
Deoxyribonucleic Acid Transcription and Protein Synthesis (Box 1.19)
The principle key factor of protein synthesis remains in the DNA sequence of the nucleus. At first the portion of DNA template is copied into the messenger RNA (mRNA).23
This mRNA is processed within the nucleus and finally comes out from the nucleus to the cytoplasm through nuclear pore. Within the cytoplasm on the small subunit of ribosome attached with RER the mRNA is decoded and protein is synthesized with the help of t-RNA.
The two major steps of the protein synthesis are:
- Transcription: It is the process of making mRNA (messenger RNA) from DNA.
- Translation: The process of synthesizing protein from the mRNA code is known as translation.
Transcription (Fig. 1.22)
In this process, m-RNA is formed from the particular sequence of nucleotides of DNA carrying the genetic information to make a particular protein. Therefore, in this step the information of DNA is transferred to the corresponding m-RNA. The basic information of DNA remains same so the process is known as transcription. RNA is essentially same as DNA in structure except in certain points: (a) This is a liner polymer of nucleotides, (b) here the sugar moiety is ribose, and (c) RNA contains the base uracil (U) instead of thymine (T).
There are three important steps of transcription: (a) initiation, (b) elongation, and (c) termination.
- Initiation: Initiation of the m-RNA synthesis needs a DNA chain, transcription factors and RNA polymerase II. At first TATA binding protein (TBP) binds with TATA box of DNA. TBP is a part of general transcription factor called TFIID. The binding of TFIID promotes the binding of another protein known as TFIIB. This complex helps in binding RNA polymerase to DNA. After the recruitment of RNA polymerase another two transcription factors TFIIE and TFIIH binds with this complex and initiation complex is completed.
- Elongation: The helicases enzyme of TFIIH unwinds DNA and the RNA polymerase starts synthesis of m-RNA from the DNA strand. RNA polymerase now moves from the promoter and elongation phase starts. The DNA strand is read from 3’ to 5’ direction and subsequently RNA strand is made from 5’ to 3’ direction.
- Termination: When the RNA polymerase reaches to the terminal codon of DNA the m-RNA synthesis is stopped.
Fig. 1.22: Schematic diagram showing mRNA transcription from DNA. Intron regions are sliced out and ultimately mRNA is formed from pre-mRNA.
Translation (Fig. 1.23)
In translation phase, “information” of m-RNA is decoded and the protein is synthesized. The newly formed mRNA passes through the nuclear pore and binds properly with the small unit of ribosome. Ribosome contains many proteins and ribosomal RNA (rRNA). There are four nucleotides (adenine, guanine, cytosine and uracil) in RNA and each group of three consecutive nucleotides of m-RNA is called a “codon” (Box 1.20). Each codon indicates one specific amino acid. More than one codon may also specify a particular amino acid. Now for each codon of m-RNA there is a specific (transfer RNA) tRNA carrying the complementary codon nucleotide sequence called “anticodon”. Each such tRNA molecule is linked with a particular amino acid. The recognition and attachment of the specific amino acid with this tRNA is dependent on enzymes known as aminoacyl-tRNA synthetase.24
Fig. 1.23: Translation of mRNA to protein synthesis in ribosomal surface is highlighted. Information of mRNA is decoded and tRNA with complementary anti-codon brings a specific amino acid to form a protein.
At first mRNA is attached with the surface of ribosome. Then specific tRNA is attached with the start codon of the mRNA carrying a particular amino acid. Ribosome moves from 5’ to 3’ direction of mRNA and then a new tRNA with another particular amino acid is attached next to the previous one. The former tRNA is released from the mRNA. This process continues till the end of the tRNA meets the stop codon and ribosome stops translation. The complete protein is synthesized and comes to the cytoplasm.
REFERENCES
- Singer SJ, Nicolson GL. The Fluid Mosaic Model of the Structure of Cell Membranes. Science. 1972;175(4023):720–31.
- Tuteja N. Signaling through G protein coupled receptors. Plant Signal Behav. 2009;4(10):942–7.
- Macara IG. Parsing the polarity code. Nature Rev Mol Cell Biol. 2004;5:220–31.
- Martin-Belmonte F, Perez-Moreno M. Epithelial cell polarity, stem cells and cancer. Nat Rev Cancer. 2012;12:23–38.
- Hugo H, Ackland ML, Blick T, et al. Epithelial-mesenchymal and mesenchymal-Epithelial transitions in carcinoma progression. J Cell Physiol. 2007;213:374–83.
- Wu Y, Zhou BP. New insights of epithelial-mesenchymal transition in cancer metastasis. Acta Biochim Biophys Sin. 2008;40:643–50.
- Royer C, Lu X. Epithelial cell polarity: a major gatekeeper against cancer? Cell Death Differentiation. 2011;18:1470–7.
- Hazan RB, Kang L, Whooley BP, et al. N-cadherin promotes adhesion between invasive breast cancer cells and the stroma. Cell Adhes Commun. 1997;4(6):399–411.
- Ellgaard L, Helenius A. ER quality control: towards an understanding at the molecular level. Curr Opin Cell Biol. 2001;13(4):431–7.
- Wilson C, Venditti R, Rega LR, et al. The Golgi apparatus: an organelle with multiple complex functions. Biochem J. 2011;433(1):1–9.
- Glick BS, Malhotra V. The curious status of the Golgi apparatus. Cell. 1998;95:883–9.
- McBride HM, Neuspiel M, Wasiak S. Mitochondria: more than just a powerhouse. Curr Biol. 2006;16(14):R551–60.
- Harris SL, Levine AJ. The p53 pathway: positive and negative feedback loops. Oncogene. 2005;24:2899–908.
- Jurgensmeier JM, Xie Z, Deve raux Q, et al. Bax directly induces release of cytochrome c from isolated mitochondria. Proc Natl Acad Sci USA. 1998;95:4997–5002.
- Alston CL, Rocha MC, Lax NZ, et al. The genetics and pathology of mitochondrial disease. J Pathol. 2016.
- Mancuso M, Orsucci D, Angelini C, et al. Redefining phenotypes associated with mitochondrial DNA single deletion. J Neurol. 2015;262:1301–9.
- Fuchs E, Weber K. Intermediate filaments: Structure, dynamics, function, and disease. Annu Rev Biochem. 1994;63:345–82.
- Foisner R, Gerace L. Integral membrane proteins of the nuclear envelope interact with lamins and chromosomes, and binding is modulated by mitotic phosphorylation. Cell. 1993;73:1267–79.
- Dey P. Nuclear margin irregularity and cancer: a review. Anal Quant Cytol Histol. 2009;31(5):345–52.
- Beck M, Lucic V, Forster F, et al. Snapshots of nuclear pore complexes in action captured by cryoelectron tomography. Nature. 2007;449:611–5.
- Tsutsui KM, Sano K, Tsutsui K. Dynamic view of the nuclear matrix. Acta Med Okayama. 2005;59(4):113–20.
- Jenuwein T, Allis CD. Translating the histone code. Science. 2001;293(5532):1074–80.
- Boisvert FM, Koningsbruggen SV, Navascues J, et al. The multifunctional nucleolus. Nat Rev Mol Cell Biol. 2007;8:574–85.
- Sirri V, Roussel P, Hernandez-Verdun D. The AgNOR proteins: qualitative and quantitative changes during the cell cycle. Micron. 2000;31:121–6.
- Watson JD, Crick FHC. A structure for deoxyribose nucleic acid. Nature. 1953;171(4356):737–8.
- An integrated encyclopedia of DNA elements in the human genome. ENCODE Project Consortium. Nature. 2012;489(7414):57–74.
- Elgar G, Vavouri T. Tuning into the signals: noncoding sequence conservation in vertebrate genomes. Trends Genet. 2008;24(7):344–52.
- Ohmori H, Friedberg EC, Fuchs RP, et al. The Y-family of DNA polymerases. Mol Cell. 2001;8(1):7–8.
- Vashee S, Cvetic C, Lu W, et al. Sequence-independent DNA binding and replication initiation by the human origin recognition complex. Genes Dev. 2003;17:1894–908.
- Bell SP, Dutta A. DNA replication in eukaryotic cells. Annu Rev Biochem. 2002;71:333–74.
Review Questions
- Increased expression of all these molecules is responsible for epithelial mesenchymal transition of epithelial cells except:
- E-cadherin
- N-cadherin
- Snail 1
- Vimentin
- The junction that connects the cell with the basal lamina:
- Adherens junction
- Tight junction
- Desmosomes
- Hemidesmosomes
- The most important site of intracellular calcium storage:
- Smooth endoplasmic reticulum
- Golgi complex
- Lysosome
- Ribosome
- During oxidative phosphorylation, a series of electron transport reactions occur in this cytoplasmic organelle:
- Smooth endoplasmic reticulum
- Golgi complex
- Mitochondria
- Ribosome
- Which one is not an intermediate filament?
- Actin
- Lamin
- Glial fibrillary acidic protein
- Desmin
- This enzyme breaks the hydrogen bonds in between the bases of DNA:
- DNA polymerase
- RNase
- Helicase
- DNA Pol I-exonuclease
- The main reactions of citric acid cycle happen in the:
- Mitochondrial matrix
- Mitochohndrial inner membrane
- Mitochohndrial outer membrane
- Golgi complex
Identify True/False
- Prokaryotic cells do not have any true nucleus and no membrane bound nucleus. (True/False)
- The loss of E-cadherin in epithelial cells enables them to detach easily and facilitates dispersion of carcinoma cells. (True/False)
- Rough endoplasmic reticulum is related with lipid synthesis. (True/False)
- Mitochondrial DNA is exclusively inherited from mother. (True/False)
- Glial fibrillary acidic protein is not an intermediate filament. (True/False)
- Euchromatin is the condensed portion of chromatin where genes are usually inactive. (True/False)
- Histone deacetylase causes chromosomal recondensation and subsequently repression of DNA transcription. (True/False)
Answers
- a. E-cadherin
- d. Hemidesmosomes
- b. Golgi complex
- c. Mitochondria
- a. Actin
- c. Helicase
- a. Mitochondrial matrix
- True
- True
- False
- True
- False
- False
- True
- False